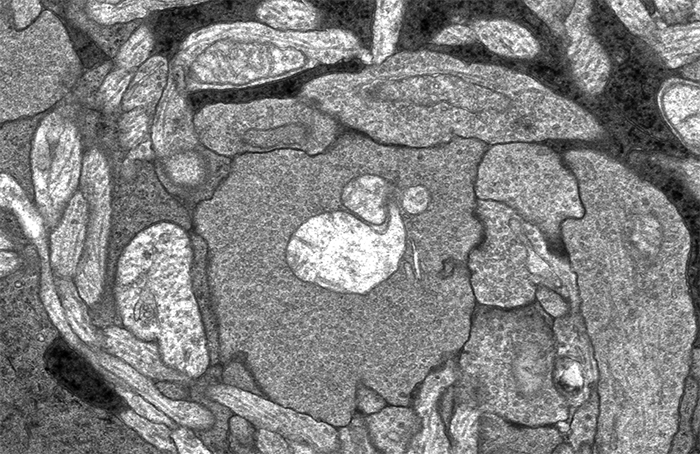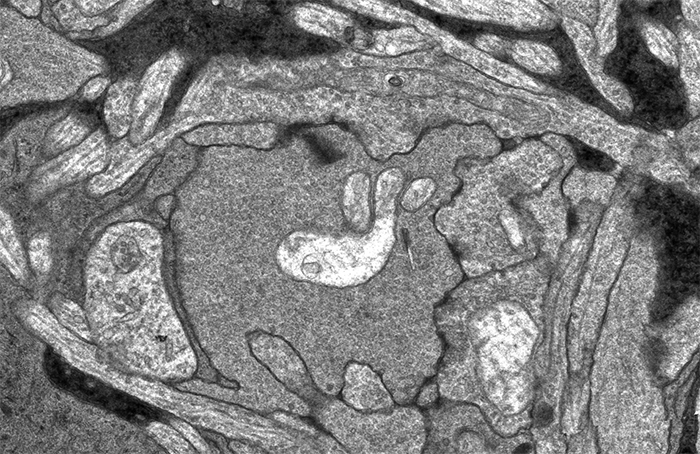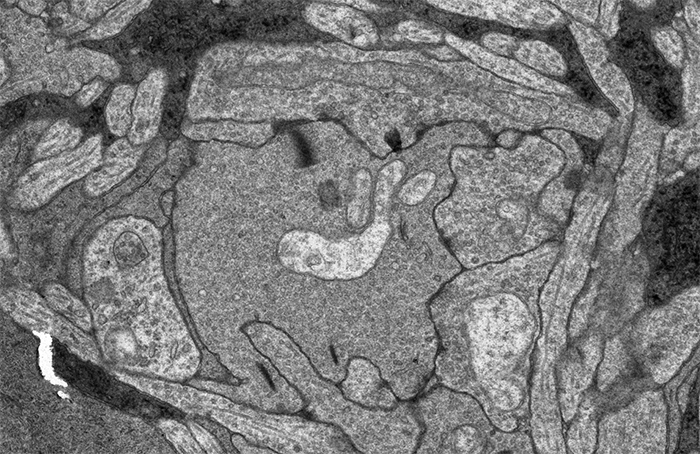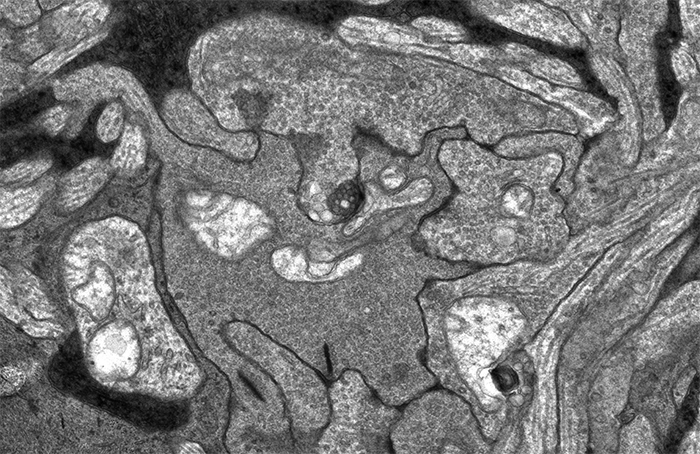
This image sequence shows a communication point between a number of cell types in your retina visualized with electron microscopy. To give you an idea of the scale of what you are looking at, the smallest diameter processes in this image are smaller than the wavelength of light and the very dark lines that appear and disappear in the image sequence are about 300 nanometers long, just greater than the wavelength of white light.
I put this sequence together after Rebecca Pfeiffer and I were looking through one of the many connectomes we’ve been assembling in the lab and I came across this bipolar cell terminal with a synaptic ribbon moving at an oblique angle (center, top 1/3rd of image). The first item of note here is that this is just a cool image sequence if you are into the ultrastructure kind of thing. The next item of note is that this is a perfect example of how difficult a computational science problem of automating image segmentation this sort of thing is. There is quite a bit going on in these image sequences which makes it a difficult computer science problem. Teaching a computer to recognize the significance of each of the objects in this field including the 3 ribbon synapses with two separate orientations, the conventional synapses, gap junctions, mitochondria, processes from horizontal cells, processes from amacrine cells and Müller cells is a monumental computer science task. But even just teaching a computer to recognize that the two ribbon synapses towards the bottom of the image and the sweeping, oblique synapse at the top of the image are the same thing is tricky.
Automating the segmentation of these types of data would be tremendously helpful as annotation is the largest rate limiting step in the analysis and visualization of these kinds of data. For now, because we don’t have automation tools, its back to work… manually segmenting these data.