This abstract was presented July 1st at the 11th International Converence of the Metabolomics Society at the University of California, Davis by Felix Vazquez-Chona, Drew Ferrell, myself and Robert E. Marc.
Metabolic dysregulation is an early hallmark of neurodegenerative diseases including Alzheimer’s disease and age-related macular degeneration. Mapping metabolic adaptation with cellular resolution and tissue- wide context is crucial to define networks regulating neuronal survival, cell death progression, and immune cell response.
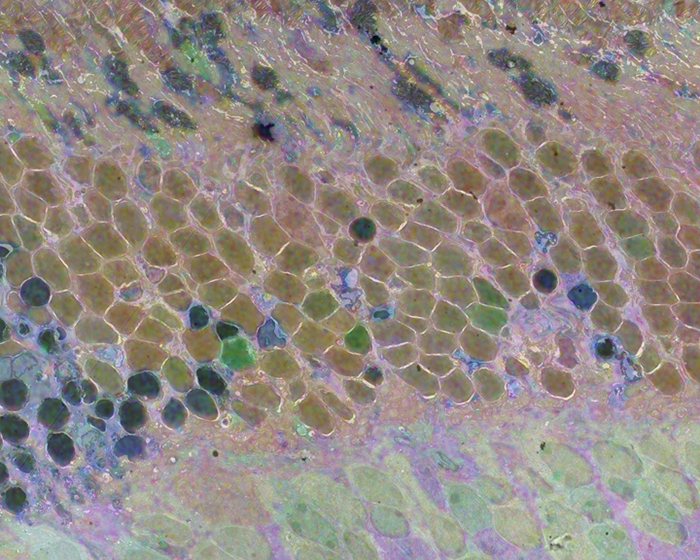
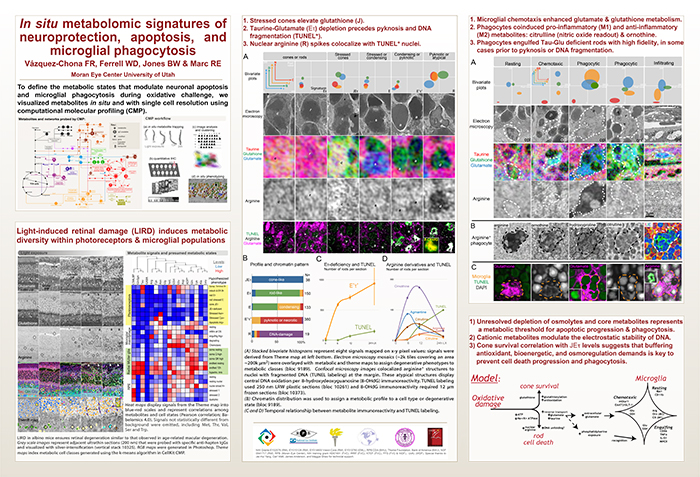
Computational Molecular Phenotyping (CMP) explores the amine metabolome (amino acids and amines). Technically, CMP metabolomics combines amine metabolite trapping, ultrathin microscopy (50-200 nm), immunodetection, pattern recognition, and clustering algorithms. Here we mapped the in situ distribution of over 30 core amine metabolites in retinal cells challenged by light-induced oxidative stress. Metabolomic profiles were phenotyped using ultrastructural, biochemical, and proteomic indices of oxidative stress.
CMP enabled precise visualization of >30 metabolites in every retinal cell. CMP resolved and phenotyped metabolomic profiles to specific degeneration and microglial functional states in the light-damaged retina. Cone photoreceptor survival correlated with enhanced antioxidant glutathione content. Rod photoreceptor apoptosis coincided with rapid depletion of organic osmolytes followed by nuclear import of cationic arginine metabolites. Delay in cell death increased necrosis and DNA damage-induced apoptosis. Microglial chemotaxis enhanced distinct signatures of glutamate and glutathione metabolism; whereas, phagocytosis coinduced classic (M1) and alternative (M2) arginine metabolites of macrophage activation.
CMP discovers and phenotypes cell classes, tracks cell state, and maps disease with single-cell resolution in any tissue or organism.
After browsing the Marclab site that you referenced, I have some questions about early brain development. The following paragraph is a quote from Wikipedia about human evolution.
“The earliest documented members of the genus Homo are Homo habilis, which evolved around 2.8 million years ago;[4] and it is arguably the earliest species for which there is positive evidence of use of stone tools. The brains of these early hominins were about the same size as that of a chimpanzee, although it has been suggested that this was the time in which the human SRGAP2 gene doubled, producing a more rapid wiring of the frontal cortex. During the next million years a process of rapid encephalization began, and with the arrival of Homo erectus in the fossil record, cranial capacity had doubled to 850 cm3.[5] This increase in human brain size is equivalent to every generation having an additional 125,000 neurons more than their parents.”
The quote states that brain size doubled in a million years to a size of 850 cm^3. That would mean the brain size started at 425 cm^3 a million years earlier. The Marclab site suggests that current brain size is about 1300 cm^3. Based on that figure my questions are as follows.
First question. Is it reasonable to assume that a brain of 425 cm^3 would have the same proportion of neurons as a larger brain? In other words, if a current brain has about 100 billion neurons in 1300 cm^3, would a primitive brain of 425 cm^3 have about 32.7 billion neurons and an 850 cm^3 brain have about 65.4 billion neurons.
If those assumptions are reasonable then there would be about 32.7 billion neurons added in a million years of evolutionary brain development. Assuming 20 years per generation for early hominids would give us 50 thousand generations. Dividing the 32.7 billion by 50 thousand would give us about 654 thousand new neurons per generation rather than the 125 thousand that the Wikipedia quote suggests.
My second question. On average, how many synapses would have to form for each new neuron that was added in brain development? I have seen figures ranging from one to ten thousand synapses per neuron.
Thanks for your help.